import mglearn
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import time
from sklearn.model_selection import train_test_split, RepeatedStratifiedKFold
from sklearn.model_selection import GridSearchCV, cross_validate
from sklearn.datasets import load_breast_cancer
from sklearn.datasets import make_moons
from sklearn.tree import DecisionTreeClassifier
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder
from sklearn.tree import export_graphviz12 Decision Trees for Machine Learning
Click here to download this chapter as a Jupyter (.ipynb) file.
In this chapter we learn how decision trees may be used for machine learning. A decision tree model may be used for both classification and regression tasks. Models based on decision trees have a few strong advantages. They are fairly easy to visualize and understand, at least for smaller trees, and there is no need to scale numeric data before fitting a tree-based model. They work well with data that has a mix of categorical and numeric features.
12.0.1 Module and Function Imports
12.1 Decision Trees
A decision tree is a hierarchy of if/else questions, leading to a decision. For example, consider a classification task in which we are trying to classify animals as various species depending on their characteristics, such as if they have feathers, if they can fly, and if they have fins. the decision tree shown below could be used to do the classification. We start at the top of the tree and ask questions as we go down. After each question the data is split into two directions, based on the answer to the question. Each instance in the data will end up at one leaf at the bottom row of the tree, and we can classify the species of that instance based on where it ends up at the bottom of the tree.
mglearn.plots.plot_animal_tree()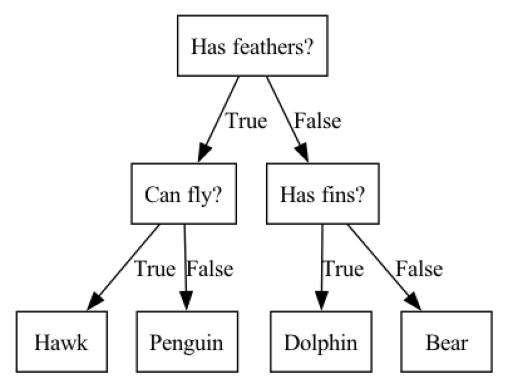
12.1.1 Some tree vocabulary
Before we continue with our exploration of tree-based models we should learn some basic vocabulary related to these models. Each point in the tree at which a splitting decision could be made is called a node. Nodes at the bottom of the tree are called leaf nodes or leaves. The tree can be described by its depth. The depth is the number of splits it takes to reach the lowest leaf node(s) from the top level
12.1.2 How does the tree determine how to split at each node?
Note that there are many specific implementations of decision trees, all of which use slightly different algorithms. The explanation provided below is a generic explanation that applies to most decision tree implementations.
The goal of a decision tree is to recursively split the dataset into subsets that are as homogeneous as possible with respect to the target variable. For example, if we start with 50 instances of data in which 25 instances have the target variable value ‘malignant’ and 25 instances have the target variable value ‘benign’, a split that resulted in all the ‘malignant’ instances on one side of the split and all the ‘benign’ instances on the other side of the split would be a perfect split, because the two leaves that resulted from the split are both homogeneous with respect to the target variable. It would be very unusual to encounter such a perfect split in a real classification task, but we can use that extreme example to understand the general concept; the more homogeneous the leaves resulting from the split are the better the split is considered to be. Another term for homogeneity is purity. At each node the tree-building algorithm finds the split that results in the greatest weighted average purity of the resulting leaves. There are several different ways that purity or impurity can be measured. Two common measures of impurity are Gini impurity and entropy. You may also encounter the term information gain which refers to a reduction in entropy.
Let’s take a closer look at Gini impurity. For binary classification, Gini impurity ranges from 0 to 0.5, where 0 represents pure nodes (all instances in the node belong to the same class), and 0.5 represents maximum impurity (an equal distribution of instances across all classes). It is calculated for a node as as 1 minus the sum of squared class proportions with the following formula, where \(k\) is the number of classes and \(p_i\) is the proportion of class \(i\) in the node: \[G = 1 - \sum_{i=1}^{k} {p_i}^2\]
To find the best split at each node the tree algorithm considers all the possible splits for each feature and selects the split that reduces impurity the most. This process continues recursively until a stopping criterion is met, such as reaching a maximum tree depth, having a minimum number of instances in a leaf node, or when further splits do not reduce impurity by more than a minimum threshold.
The decision tree-building process is essentially a greedy algorithm, making locally optimal choices at each node. Note that this might not always result in the optimal tree structure at a global level.
12.1.3 Visualization of a Tree-Based Binary Classification Model with 2 Features
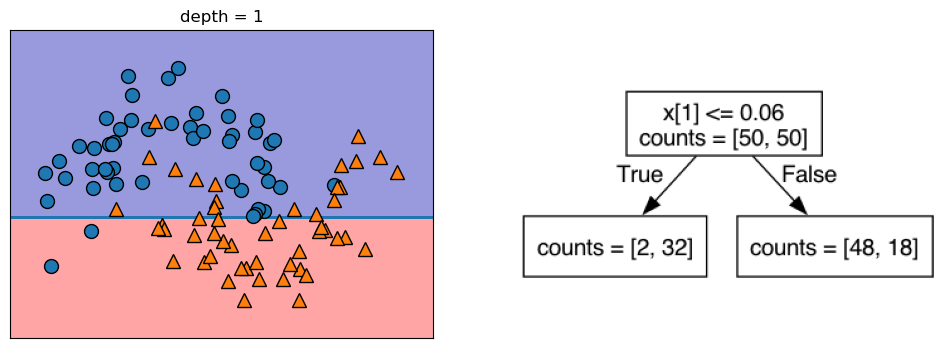
Let’s take a look at some visualizations that show what happens in the first several splits of a tree-building process. We begin by visualizing the data. There are two features, and the target variable has two classes, blue circles and orange triangles. The next few visualizations show the decision boundary created by each split and the current state of the decision tree after each split. Note how adding levels to the tree allows the tree more flexibility to fit the training data. Consider how that will affect generalization performance.
mglearn.plots.plot_tree_progressive()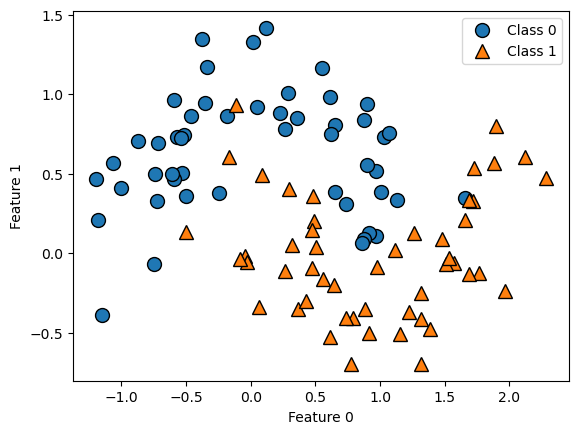
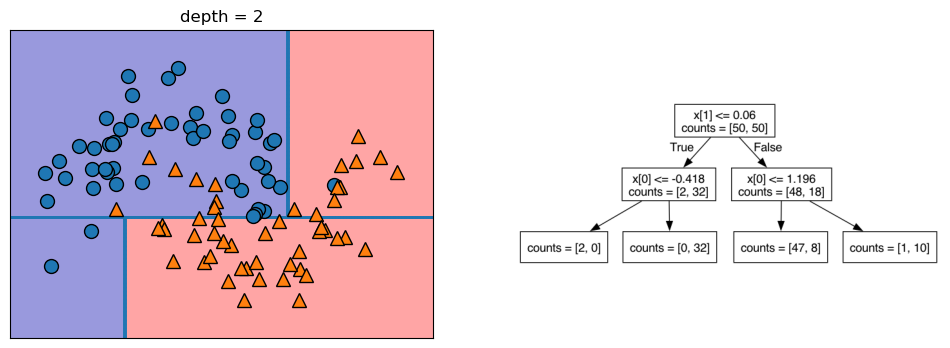
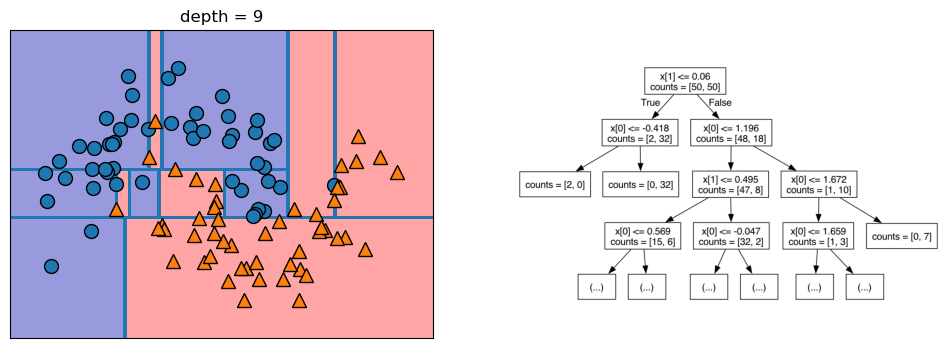
12.1.4 Controlling the Complexity of Decision Trees
If a decision tree model is permitted to continue to split leaves it attempt to keep splitting until all the leaves are pure. So, the more freedom we give the tree to continue to split leaves and grow, the more flexibility we are giving it to fit the training data. We can control the flexibility of decision trees with pre-pruning or post-pruning. Pre-pruning refers to setting some parameter that causes the tree to stop growing at some pre-defined threshold. Following are some commonly-used pre-pruning parameters (see the scikit-learn documentation for others): * max_depth - maximum number of levels for the tree * max_leaf_nodes - maximum number of leaf nodes * min_samples_leaf - stops the splitting when either of the leaves resulting from the split would have fewer than this number of instances in it * min_samples_split - will not split a leaf with fewer than this number of instances in it
When we build single-tree models with scikit-learn we will vary one or more of these parameters to modify the model’s flexibility to fit the training data.
There are also approaches that utilize post-pruning. With post-pruning approaches the full tree is built and then some nodes are removed or collapsed according to some criterion. We will not be covering post-pruning approaches in this chapter.
The primary purpose of both pre-pruning and post-pruning is to limit the complexity (flexibility) of the tree so that it does not overfit the training data.
12.1.5 Estimate Generalization Performance for and Tune the Decision Tree model
Let’s run through the full model-building process for the decision tree algorithm on the Titanic data, tuning the min_samples_leaf parameter:
- set up a grid search
- run the grid search once to see if parameter range selected is reasonable
- run a nested cross validation to estimate generalization performance. Recall that a nested cross validation is a cross validation that uses a grid search as the estimator. It is considered the best way to estimate a model’s generalization performance. Note that it cannot be used to tune the parameters of the final model, however, because each of the interior grid searches may arrive at different settings for the parameters. Because of this, a nested cross validation is typically used to select the best algorithm for the model (tree, logistic regression, K-nn, etc.) and then a separate grid search is performed on the best algorithm using all the data to set the final model parameters
- perform final grid search on all the data to tune the model
12.1.6 Load the Titanic Data
In the cell below we will import a subset of the Titanic data, which describes passengers on the Titanic. Below is a brief description of the variables: * Survived - the target variable. Has values 0 or 1, with 1 representing that the passenger survived the disaster * Pclass - represents class of ticket * Sex - represents the sex of the passenger * Age - represents the age of the passenger * SibSp - represents number of siblings or spouse traveling with the passenger * Parch - represents the number of parents or children traveling with the passenger * Embarked - represents the port from which the passenger embarked.
Note: This is only a subset of the actual Titanic dataset.
titanic = pd.read_csv('https://neuronjolt.com/data/titanic.csv')titanic.info()<class 'pandas.core.frame.DataFrame'>
RangeIndex: 712 entries, 0 to 711
Data columns (total 7 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 Survived 712 non-null int64
1 Pclass 712 non-null int64
2 Sex 712 non-null object
3 Age 712 non-null float64
4 SibSp 712 non-null int64
5 Parch 712 non-null int64
6 Embarked 712 non-null object
dtypes: float64(1), int64(4), object(2)
memory usage: 39.1+ KBX = titanic.drop(columns = 'Survived')
y = titanic['Survived']12.1.7 Define the grid search
The Titanic dataset has both numeric and categorical features. We don’t need to scale the numeric features, as we do with K-nn or linear models, but we do need to one-hot encode the categorical features. So, we will build a column transformer to one-hot encode the categorical features. Note that if we don’t specify an operation to perform on the numeric columns they will be dropped from the analysis unless we explicitly tell the column transformer to pass them through unchanged. We can do that by setting the remainder parameter to 'passthrough' within the ColumnTransformer() function.
Since there are not many rows of data, we will use 3 x stratified 10-fold cross validation as our splitting strategy. We set the n_jobs parameter to -1, which tells scikit learn to parallelize the grid search over all available processor cores.
# Define column transformer to one-hot encode the categorical features
# and pass through the numeric features
ct_dt = ColumnTransformer([
('ohe', OneHotEncoder(handle_unknown = 'ignore'), ['Sex', 'Pclass', 'Embarked'])
], remainder = 'passthrough')
# Combine the column transformer with the estimator in a pipeline
pipe_dt = Pipeline([
('pre', ct_dt),
('dt', DecisionTreeClassifier())
])
# Define the parameter grid
pgrid_dt = {'dt__min_samples_leaf': range(1, 102, 5)}
# Define a custom splitter to do 3 x stratified 10-fold cv
rskf_3x10 = RepeatedStratifiedKFold(n_splits = 10, n_repeats = 3)
# Define the grid search
gs_dt = GridSearchCV(estimator = pipe_dt,
param_grid = pgrid_dt,
cv = rskf_3x10,
n_jobs = -1,
return_train_score = True)12.1.8 Run the grid search once to check parameter range
In the initial grid search we are varying the parameter min_samples_leaf from 1 to 101 by 5s. This parameter controls how far the tree can keep splitting, because a split will not be done if it would result in a leaf with fewer instances in it than the min_samples_leaf setting. The choice of 1 to 101 was made to get a fairly wide range, but go by 5s so the initial grid search doesn’t take too long. We will run the grid search once to see if this parameter range is appropriate. For the grid search we use in the nested cross validation below we want to use a parameter range centered around the best parameter setting from this initial grid search, but if the best parameter setting from the initial grid search is at the high end of the range we should extend the range and try again.
gs_dt.fit(X, y)GridSearchCV(cv=RepeatedStratifiedKFold(n_repeats=3, n_splits=10, random_state=None),
estimator=Pipeline(steps=[('pre',
ColumnTransformer(remainder='passthrough',
transformers=[('ohe',
OneHotEncoder(handle_unknown='ignore'),
['Sex',
'Pclass',
'Embarked'])])),
('dt', DecisionTreeClassifier())]),
n_jobs=-1, param_grid={'dt__min_samples_leaf': range(1, 102, 5)},
return_train_score=True)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
Parameters
['Sex', 'Pclass', 'Embarked']
Parameters
['Age', 'SibSp', 'Parch']
passthrough
Parameters
gs_dt.best_params_{'dt__min_samples_leaf': 6}We used the best_params_ property of the fitted grid search object to see what the grid search identified as the best parameter setting for min_samples_leaf. It found that 6 was the best parameter setting. This suggests that our range of 1 to 101 might be a bit wide, because the best parameter setting found on the initial grid search was near the low end of that range. However, before we proceed to the nested cross validation to estimate generalization performance for the decision tree model on the breast cancer data we will create a validation curve plot to see the pattern of how test performance varies with changes in the min_samples_leaf parameter.
# Put the results from the grid search into a DataFrame
results = pd.DataFrame(gs_dt.cv_results_)# Make the validation curve plot
plt.plot(results['param_dt__min_samples_leaf'],
results['mean_train_score'],
label="training accuracy")
plt.plot(results['param_dt__min_samples_leaf'],
results['mean_test_score'],
label="test accuracy")
plt.axhline(max(results['mean_test_score']), linestyle='dotted', color='gray')
plt.axvline(results['param_dt__min_samples_leaf'].iloc[results['mean_test_score'].idxmax()],
linestyle='dotted', color='gray')
plt.title("Validation Curve Graph for Decision Tree")
plt.ylabel("Prediction Accuracy")
plt.xlabel("Min Samples per Leaf")
plt.xlim(max(results['param_dt__min_samples_leaf']), min(results['param_dt__min_samples_leaf']))
plt.legend();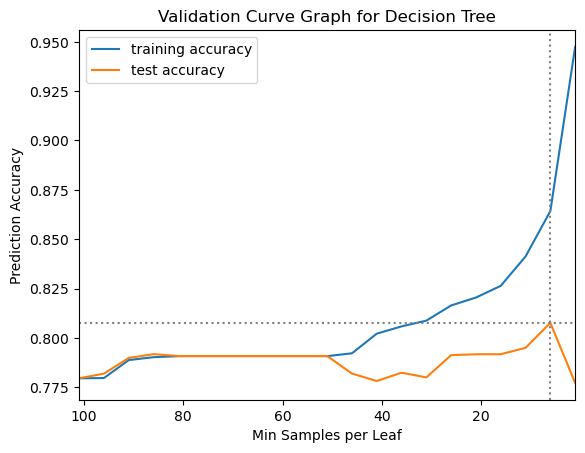
12.1.9 Revise Parameter Grid and Re-Define Grid Search before Nested Cross Validation
We can see from the validation curve plot that the mean test performance from the grid search has its peak at a min_samples_leaf parameter setting of 6 and then declines sharply. To help the nested cross validation run faster we will narrow the range of the parameter grid before running the nested cross validation. Having fewer parameter settings for each grid search to check will take less time, but we don’t want to narrow the range too much, though, because we want to make sure we find the best parameter setting at each split. We will also not save training scores for the nested cross validation. This will also help the grid search to run a bit faster. When we change the parameter grid we need to re-define the grid search as well, to pull in the new parameter grid.
# Re-define the parameter grid
pgrid_dt = {'dt__min_samples_leaf': range(1, 16)}
# Re-define the grid search with the narrowed parameter grid
gs_dt = GridSearchCV(estimator = pipe_dt,
param_grid = pgrid_dt,
cv = rskf_3x10,
n_jobs = -1)12.1.10 Perform the Nested Cross Validation: Use the grid search as the estimator in a cross validation
For the nested cross validation we will use stratified 10-fold cross validation. Note that we do not set the n_jobs parameter within the cross_validate function below, because it is already set in the grid search that serves as the estimator in the cross_validate function. It is recommended to only set the n_jobs parameter in one level of a nested grid search, either in the grid search or the cross validate but not both.
# Perform the nested cross validation
results_dt = cross_validate(estimator = gs_dt,
X = X,
y = y,
cv = 10)12.1.10.1 Calculate the mean test score as estimate of generalization performance
Below we calculate the mean score on the test folds from the nested cross validation. This is our best estimate of the generalization performance of the decision tree model on the breast cancer data. It is also useful to take a look at the standard deviation of the test score to see how sensitive the test scores are to the data splits.
results_dt['test_score'].mean()np.float64(0.7894757433489827)results_dt['test_score'].std()np.float64(0.05903215268777309)So, our estimate of the generalization performance of the decision tree model on the Titanic data is around 79% accuracy, with a standard deviation of around 6%.
In a full model building process we would perform nested cross validation for each algorithm we were considering (K-nn, logistic regression, decision tree, etc.) to determine which algorithm has the best estimated generalization performance on our data. Then, we would re-run the grid search for the best algorithm on the unsplit data to tune it and prepare it to make future predictions.
final_model = gs_dt.fit(X, y)If we had new data, we would use this final model’s predict method to make predictions of the target variable, with code such as the following:
preds = final_model.predict(new_data)